
Discovery on How CRISPR Proteins Identify Targets
July 26, 2017| |
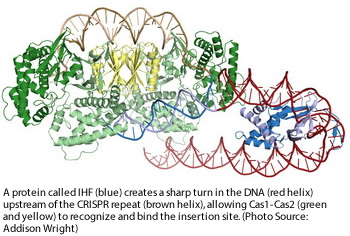 Researchers at University of California, Berkeley (UC Berkeley) have discovered how Cas1-Cas2, the proteins that allow the CRISPR immune system in bacteria to adapt to new viral infections, identify the site in the genome where they insert viral DNA so they can recognize it later and mount an attack. In the July 20 paper published in Science, Jennifer Doudna and her group reports capturing structures of Cas1-Cas2 in the act of inserting viral DNA into the CRISPR region. The structures revealed that a third protein, IHF, binds near the insertion site and bends the DNA into a U-shape, allowing Cas1-Cas2 to bind both parts of the DNA simultaneously. The team also discovered that the reaction requires that the target DNA bend and partly unwind, something that occurs only at the proper target.
Researchers at University of California, Berkeley (UC Berkeley) have discovered how Cas1-Cas2, the proteins that allow the CRISPR immune system in bacteria to adapt to new viral infections, identify the site in the genome where they insert viral DNA so they can recognize it later and mount an attack. In the July 20 paper published in Science, Jennifer Doudna and her group reports capturing structures of Cas1-Cas2 in the act of inserting viral DNA into the CRISPR region. The structures revealed that a third protein, IHF, binds near the insertion site and bends the DNA into a U-shape, allowing Cas1-Cas2 to bind both parts of the DNA simultaneously. The team also discovered that the reaction requires that the target DNA bend and partly unwind, something that occurs only at the proper target.
CRISPR, the unique region of DNA where snippets of viral DNA are stored for future reference, allows the cell to recognize any virus that tries to re-infect. The viral DNA alternates with the "short palindromic repeats," which serve as the recognition signal to direct Cas1-Cas2 to add new viral sequences.
Specific recognition of these repeats by Cas1-Cas2 restricts integration of viral DNA to the CRISPR array, allowing it to be used for immunity and avoiding the potentially fatal effects of inserting viral DNA in the wrong place. This research opens the door for modification of the proteins themselves. Tweaking the proteins means that researchers might be able to redirect them to sequences other than the CRISPR repeat and expand their application into organisms without their own CRISPR locus.
For more details, read the UC Berkeley News.
| |
Biotech Updates is a weekly newsletter of ISAAA, a not-for-profit organization. It is distributed for free to over 22,000 subscribers worldwide to inform them about the key developments in biosciences, especially in biotechnology. Your support will help us in our mission to feed the world with knowledge. You can help by donating as little as $10.
-
See more articles:
-
News from Around the World
- Web Portal Established to Speed up Genetic Research in Plants
- India's B.R. Barwale Passes Away
- African Biosafety Regulators Embrace Biosafety Communication at ABBC 2017
- Biotechnology Praised at the Biggest Agri Expo in Uganda
- Adoption of GE Crops in the US
- New Gene in Corn Confers Resistance to Multiple Diseases
- Study Finds Plants Reprogram Their Genetic Material to Fight Pathogens
- China Gives Import Approval for Corn Rootworm Resistant Trait
- Russia Drafts Regulations for GE Feed
-
Research Highlights
- Overexpression of BoC3H Gene Enhances Salt Stress Tolerance in Broccoli
- Study on the Effect of Light and Temperature on CBF14 Expression in Wheat and Barley
-
Beyond Crop Biotech
- NAS Announces Awards for Science Communication
- Researchers Develop Plant-produced Vaccine Candidates Against Bluetongue Virus
-
Plant
- CRISPR-Cas9 and CRISPR-Cpf1 Mediated Genome Editing of EPFL9 Gene in Rice
- Discovery on How CRISPR Proteins Identify Targets
-
Read the latest: - Biotech Updates (April 22, 2026)
- Gene Editing Supplement (April 29, 2026)
- Gene Drive Supplement (February 22, 2023)
-
Subscribe to BU: - Share
- Tweet
